- About
-
Research
- Agronomy and farming systems
-
Agricultural crop research
-
Research projects - agriculture
- About SASSA-SAI
- CTP for Sustainable Agricultural Innovation
- Climate Ready Beans - workshop presentations (March 2022)
- Crop diversity HPC cluster
- Final project workshop
- Get involved
- List of materials
- News and updates
- Partners
- The research team
- UK Cereal Pathogen Virulence Survey
- UK wheat varieties pedigree
- Weed management - IWM Praise
- Crop breeding
- Crop characterisation
- Data sciences
- Genetics and pre-breeding
- Plant biotechnology
- Plant pathology and entomology
- Resources
-
Research projects - agriculture
- Horticultural crop research
- Crop Science Centre
- Research Projects
- Research Publications
-
Services
- Analytical Services
- Business Development
- Commercial trial services
- Membership
- Plant breeding
- Plant characterisation
- Seed certification
-
Training
-
Technical agronomy training
- Advanced crop management of bulb onions
- Advanced crop management of vegetable brassicas
- Advanced nutrient management for combinable crops
- Benefits of cover crops in arable systems
- Best practice agronomy for cereals and oilseed rape
- Developing a Successful Strategy for Spring Crops
- Disease Management and Control in Cereal Crops
- Incorporating SFI options into your rotation
- Protected Environment Horticulture – Best Practice
- Techniques for better pest management in combinable crops
- Crop inspector and seed certification
- Licensed seed sampling
-
Technical agronomy training
- News & Views
- Events
-
Knowledge Hub
- Alternative and break crops
-
Crop genetics
- POSTER: Diversity enriched wheat (2025)
- POSTER: Genetics of wheat flag leaf size (2024)
- POSTER: Wheat yield stability (2024)
- Poster: Traits for future cereal crops (2022)
- POSTER: wild wheat fragment lines (2022)
- POSTER: Improving phenotyping in crop research (2022)
- PRESENTATION: Plant breeding for regen ag
- Poster: Designing Future Wheat (2020)
- Crop nutrition
-
Crop protection
- POSTER: Understanding the hierarchy of black-grass control (2025)
- POSTER: Emerging weed threats (2025)
- POSTER: Disease control in barley (2025)
- Poster: Weed seed predation in regen-ag (2024)
- POSTER: Disease control in winter wheat (2025)
- POSTER: Mode of action (2023)
- POSTER: Inter-row cultivation for black-grass control (2022)
- POSTER: UKCPVS winter wheat yellow rust in spring 2025 (2025)
- Poster: Management of Italian ryegrass (2021)
- POSTER: UKCPVS winter wheat rusts - 2024/25 review (2025)
- POSTER: UKCPVS disease monitoring and the benefit to UK growers (2025)
- POSTER: Diagnosing and scoring crop disease using AI (2025)
- POSTER: Finding new sources of Septoria resistance (2024)
- POSTER: Fungicide resistance research (2024)
- POSTER: Detecting air-borne pathogens (2024)
- POSTER: Oilseed rape diseases (2024)
- POSTER: Fungicide resistance research (2024)
- POSTER: Improving chocolate spot resistance (2022)
- Poster: Pathogen diagnostics (2022)
- Fruit
- Regen-ag & sustainability
-
Seed certification
- POSTER: Wheat DUS (2024)
- POSTER: Innovation in variety testing (2024)
- POSTER: AI and molecular markers for soft fruit (2024)
- POSTER: Barley crop identification (2023)
- POSTER: Herbage grass crop identification (2023)
- POSTER: Herbage legume crop identification (2024)
- POSTER: Minor cereal crop inspecting (2023)
- POSTER: Pulse crop identification (2023)
- POSTER: Wheat crop identification (2023)
-
Soils and farming systems
- POSTER: Checking soil health - across space and time (2024)
- POSTER: Checking soil health - step by step (2024)
- POSTERS: Changing soil management practices (2022)
- Poster: Monitoring natural enemies & pollinators (2021)
- POSTER: Soil structure and organic matter (2024)
- POSTER: Novel wheat genotypes for regen-ag (2024)
- Video: New Farming Systems project (2021)
- Video: Saxmundham Experimental Site (2021)
- POSTER: Impact of prolonged rainfall on soil structure (2024)
- POSTER: Soil & agronomic monitoring study (2024)
- POSTER: The impact of rotations & cultivations (2024)
- VIDEO: Great Soils; soil sampling guidelines (2020)
- Poster: Soil invertebrates within arable rotations (2024)
- VIDEO: Soil health assessment (2021)
- POSTER: Saxmundham - modern P management learnings
- POSTER: Saxmundham - 125 years of phosphorus management
- Poster: Soil phosphorus - availability, uptake and management (2025)
- POSTER: Morley long term experiments (2025)
- POSTER: Exploiting novel wheat genotypes for regen-ag (2025)
- Video: Saxmundham Experimental Site (2021)
- Varieties
Community Resource for Wheat and Rice Transformation - Information and Online Application Form
APPLICATION FORM
The closing date for the current round of applications is
Friday 27th January 2023
The Community Resource for Wheat and Rice Transformation provides capacity for 100 genes to be transformed into wheat and rice free of charge.
NIAB envisages that 75 genes will be transformed into wheat and 25 genes into rice. In the event that wheat or rice transformation is not fully subscribed, additional slots will be made available in the other species. We will encourage researchers working with genes from model crops to test their hypotheses in wheat and rice as appropriate to the gene studied. This will enable new genes to be evaluated more quickly in crops essential to food security worldwide, and allow researchers to amass crucial data which can be used as a basis to seek follow-on funding.
This project also provides funded capacity to characterise 50 regulatory elements in wheat and rice. Promoter and terminator sequences will be included for their expression in a wide range of tissue types, biotic and abiotic stress induced elements will also be included. Researchers can either nominate promoters to be included in this project, or provide constructs which are compatible with our vectors.
Please contact the croptransformation [at] niab.com (NIAB Crop Transformation Group) for further details on this aspect of the project.
This project has been funded for five years by the BBSRC Bioinformatics & Biological Resources fund.
Who can apply?
Academic researchers from UK who are eligible for BBSRC funding are invited to apply for the transformation of their nominated genes. Researchers may apply for several different genes, or for over-expression and silencing by RNAi of the same gene target.
Only 1 gene/construct can be included per application, however a researcher may submit up to two applications per round. Further applications will not be considered by the Advisory Group until outstanding construct(s) from the same researcher, accepted in previous rounds, have been received.
This is a resource for researchers who would like to express a gene in wheat or rice, but do not have the funding to support the transformation. There must be a clear relevance to BBSRC food security targets.
Vectors
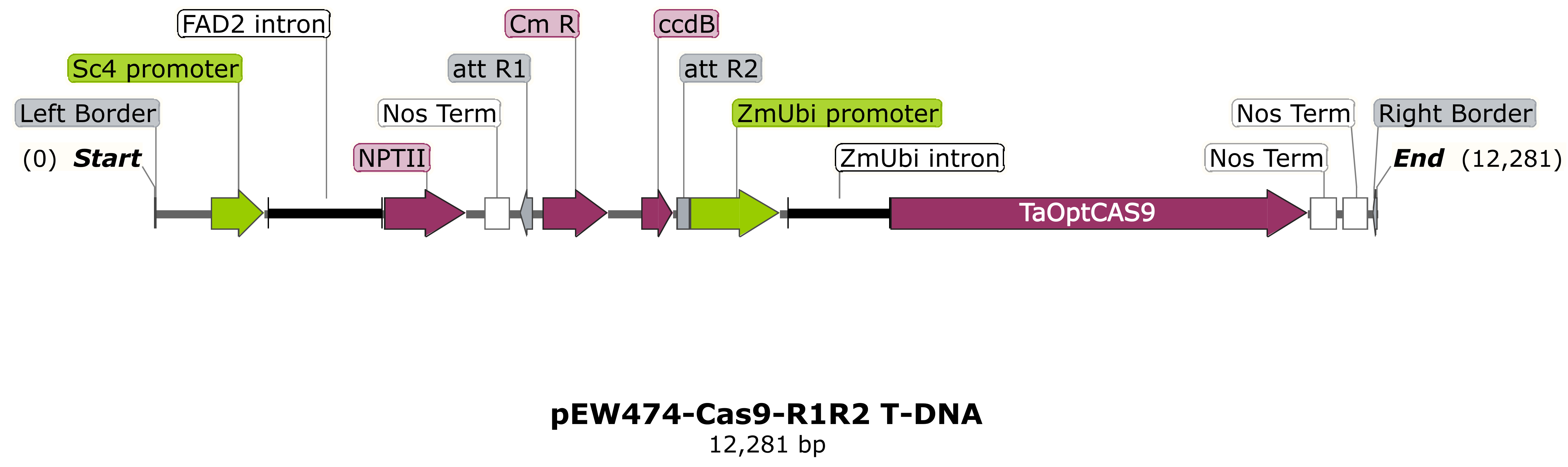
NIAB would prefer genes to be supplied in an intermediate pEntr-type construct flanked by attL1/attL2 sequences, ready for transfer into an appropriate NIAB binary vector with a selection cassette via Gateway™ cloning. Some examples of typical destination vector T-DNA regions for cereal transformation are shown below. Advice on construct design can be provided. Confirmation by sequencing that the supplied Entry-type intermediate construct is correct will be required at the construct submission stage.
NIAB typically uses either the maize ubiquitin promoter or the rice actin promoter for constitutive over-expression (pEW472-Ubi-R1R2 or pSc4Act-R1R2).
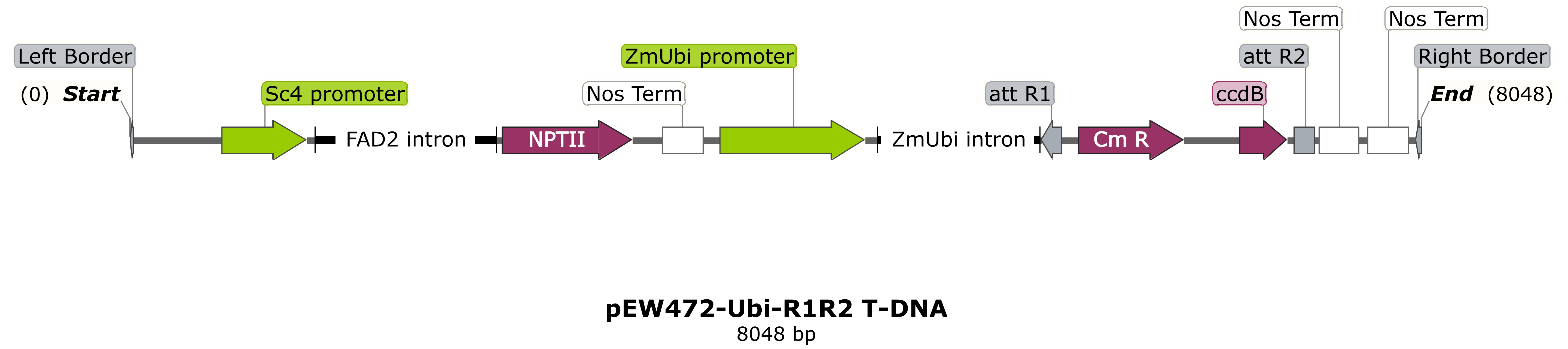
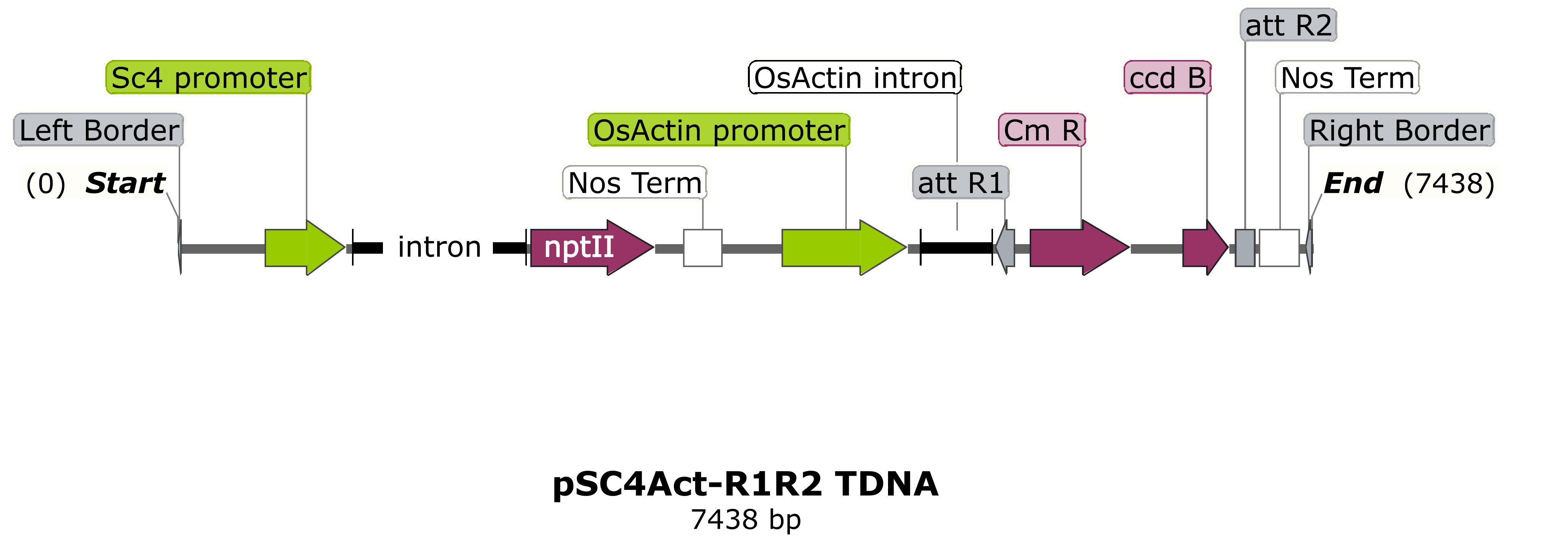 Alternatively, researchers may need to introduce their own "promoter-gene of interest-terminator" cassette, for example with the gene of interest expressed from the native sequence or an alternative promoter, (pRLF12R1R2).
Alternatively, researchers may need to introduce their own "promoter-gene of interest-terminator" cassette, for example with the gene of interest expressed from the native sequence or an alternative promoter, (pRLF12R1R2).
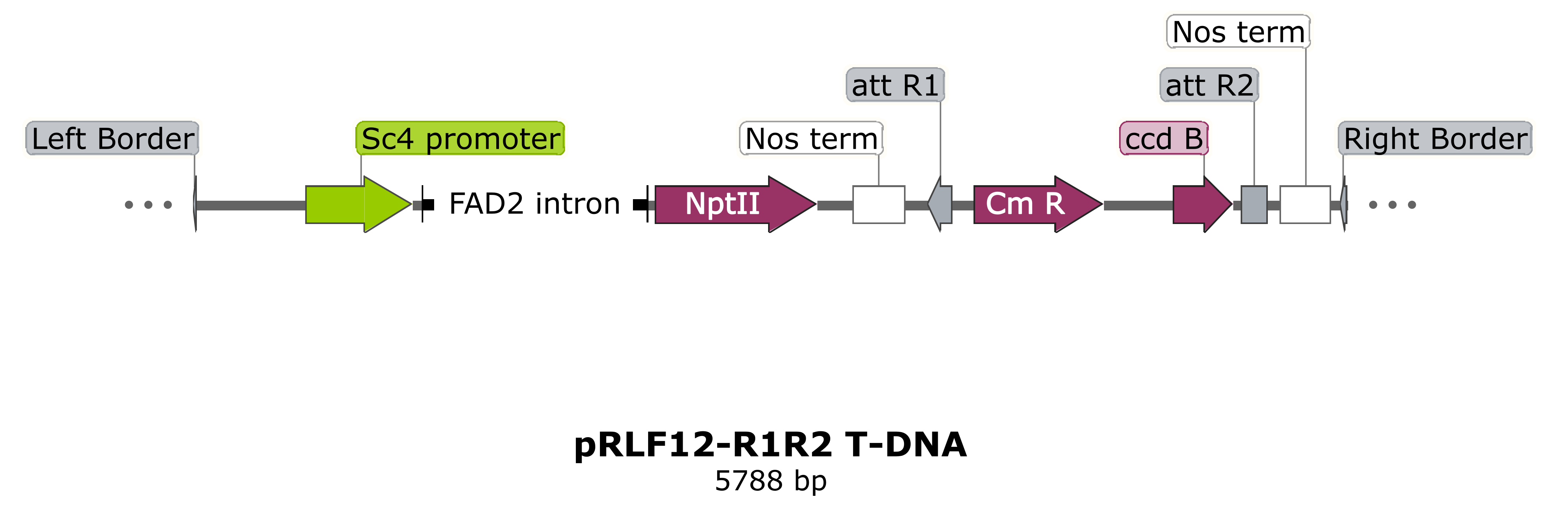 Gateway vectors for RNAi silencing or CRISPR/Cas9 editing of the target gene of interest are also available. Again, RNAi trigger or sgRNA guides should be flanked by attL1/attL2 sequences.
Gateway vectors for RNAi silencing or CRISPR/Cas9 editing of the target gene of interest are also available. Again, RNAi trigger or sgRNA guides should be flanked by attL1/attL2 sequences.
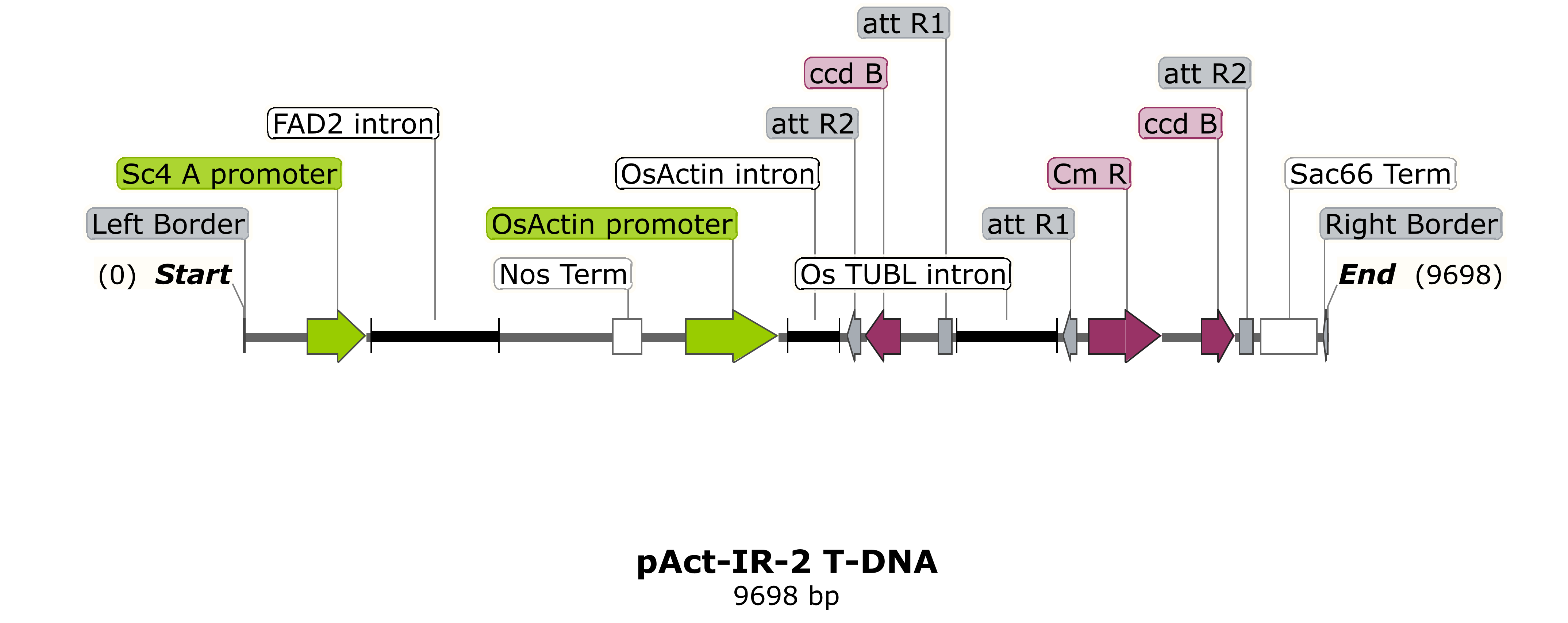
 Other vectors and promoters for traditional cloning approaches or Gateway® cloning may also be available. Goldengate constructs can also be accepted if the gene cassettes are flanked by attL sites.
Other vectors and promoters for traditional cloning approaches or Gateway® cloning may also be available. Goldengate constructs can also be accepted if the gene cassettes are flanked by attL sites.
Please contact the croptransformation [at] niab.com (NIAB Crop Transformation Group) at an early stage to discuss your cloning strategy.
Delivery
Our standard service is to deliver 30 independent transformed T0 wheat plants per construct, as plantlets at 12 weeks from experiment start to the researcher to grow in their own facilities. Non-transformed plants will be provided as control material. In the majority of cases we will use a US soft spring hexaploid wheat, Fielder, for this activity, but if spring or winter wheat germplasm with a specific end-use or disease susceptibility is necessary for your project, please include this information in your application. An adjustment to the number of plants produced with the alternative germplasm may be made.
Rice transformation will use Nipponbare, Kitaake or Donjin germplasm, with 15-20 T0 plants per construct at about eight weeks from experiment start, provided to the researcher to grow in their own facilities.
Post transformation analysis
All plantlets will be analysed by qPCR at the small rooted plantlet stage, to verify that they are transgenic and to provide information on the number of T-DNA copies introduced. Researchers will receive small rooted T0 plants and are very strongly encouraged to undertake RNA /protein expression, metabolic or phenotypic analysis on these T0 plants to prioritise lines for further investigation. The strategy for this analysis must be included in the application, together with confirmation that the researcher has the appropriate facilities to grow the plants to maturity.
The application and review process
There will be an annual call for applications each year, until the programme is completed. If the number of applications exceeds the capacity available, applications will be reviewed by our external Advisory Group. Applications will be ranked on scientific merit/relevance to BBSRC food security, the quality of the proposal and the strategy for analysis.
All applicants will be required to send a brief annual update listing all poster and oral presentations, MSc/PhD theses and papers describing these materials to the CRWRT PI for inclusion in UKRI Research Impact Assessment, and ongoing project review by the Advisory Group.





