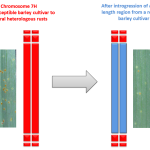
Would non-host resistance in barley prove a source of durable, race-nonspecific resistance for wheat?
Researcher / PhD Candidate : Mathilde Daniau
 Mathilde’s project consists in identifying and characterising barley genes involved in non-host resisance to leaf rust and powdery mildew both in barley and in wheat.
Mathilde’s project consists in identifying and characterising barley genes involved in non-host resisance to leaf rust and powdery mildew both in barley and in wheat.
Nonhost resistance (NHR), defined as the immunity displayed by all genotypes of a plant species towards all pathotypes of a potential pathogenic species, is the most common form of disease resistance in plants. It may be based on Pathogen-Associated Molecular Pattern (PAMP)-Triggered Immunity (PTI) and Effector-Triggered Immunity (ETI).
This durable immunity of plant genotypes to a broad-spectrum of potential pathogens suggests that NHR is highly relevant for agricultural applications. However, its genetic basis still remains unclear.
The current research project aims to a better understanding of NHR in barley through two complementary approaches:
– One, based on forward genetics, involves the fine mapping of a 80 cM QTL in barley which has shown to confer resistance to several heterologous rusts including the wheat leaf rust Puccinia triticina;
– The other one, built on reverse genetics, aims to functionally characterize two barley Receptor-Like Kinases (RLKs) genes which have shown, from previous research at IPK (Gatersleben, DE) using transient system, to play a role in NHR to the wheat powdery mildew (Blumeria graminis tritici – Bgt). The current project aims to validate this by stable over-expression studies. The transfer of these genes into wheat and the assessment of the resulting wheat transgenic lines for response to Bgt inoculation might determine whether genes involved in NHR in barley can confer host resistance when transferred to wheat.
These two approaches will lead to identify new components involved in basal resistance and to better understand nonhost resistance. This knowledge may be applied in breeding programs for resistance.
Aspects of Mathilde’s PhD research form part of a bigger, ERA-CAP project: DURESTrit (Nov. 2014-Oct. 2017), co-ordinated by Dr Patrick Schweizer, IPK, Gatersleben, Germany. Consequently Mathilde has been included in this European collaboration.
Summary of Project Findings
ESR1 consisted of two parts: (i) Fine mapping of a QTL region in the barley variety Vada that conferred resistance to four heterologous rusts; Puccinia hordei murini (Phm), P. hordei secalini (Phs), P. triticina (Pt) and P. graminis lolii (Pgl), and (ii) Cloning of two barley receptor-like kinases (RLK) that conferred NHR to the wheat-adapted pathogen, Blumeria graminis f. sp. tritici (Bgt), followed by over-expression of these barley RLKs in the wheat. Through repeated rounds of selfing recombinant plants were obtained that allowed fine mapping of three QTLs; sub-region 1 (Phm and Phs NHR), sub-region 2 (Pt NHR), and sub-region 3 (Pgl NHR). The intervals identified from previous research for Phm and Pgl could not been confirmed, therefore, subsequent work focused only of sub-region Pt. The Pt sub-region has been mapped to a genetic distance of 0.6cM, representing 20 Mb on the barley Morex genome reference. Two barley RLKs have been cloned for variety Morex and over-expressed in the wheat variety Fielder. The NHR phenotype has been confirmed in barley, variety Golden Promise, by siRNA. However, despite extensive testing, the host resistance seen in transient over-expression studies in the wheat variety Kanzler could not be duplicated in the Fielder genetic background.
SUPERVISORS:
Wageningen University: Dr. Ir. Rients E. Niks
NIAB: Dr. Lesley A. Boyd
MATHILDE’S EDUCATIONAL BACKGROUND
- 2014-2018 : PhD candidate at the Plant Breeding laboratory of Wageningen University and Research Center (NL), in partnership with the department of Genetics and Pre-Breeding at the National Institute of Agricultural Botany (NIAB, UK)
- 2011-2014 : Master’s degree in agronomy from the ISA (Lille, FR) http://www.isa-lille.com/
- 2009-2011 : Bachelor’s degree in biology and biotechnology from the Catholic University of Lille, France http://www.univ-catholille.fr/about-us/welcome.asp